News & Information
Featured

Dateline
News Resources
Podcasts and Shows

Unfold
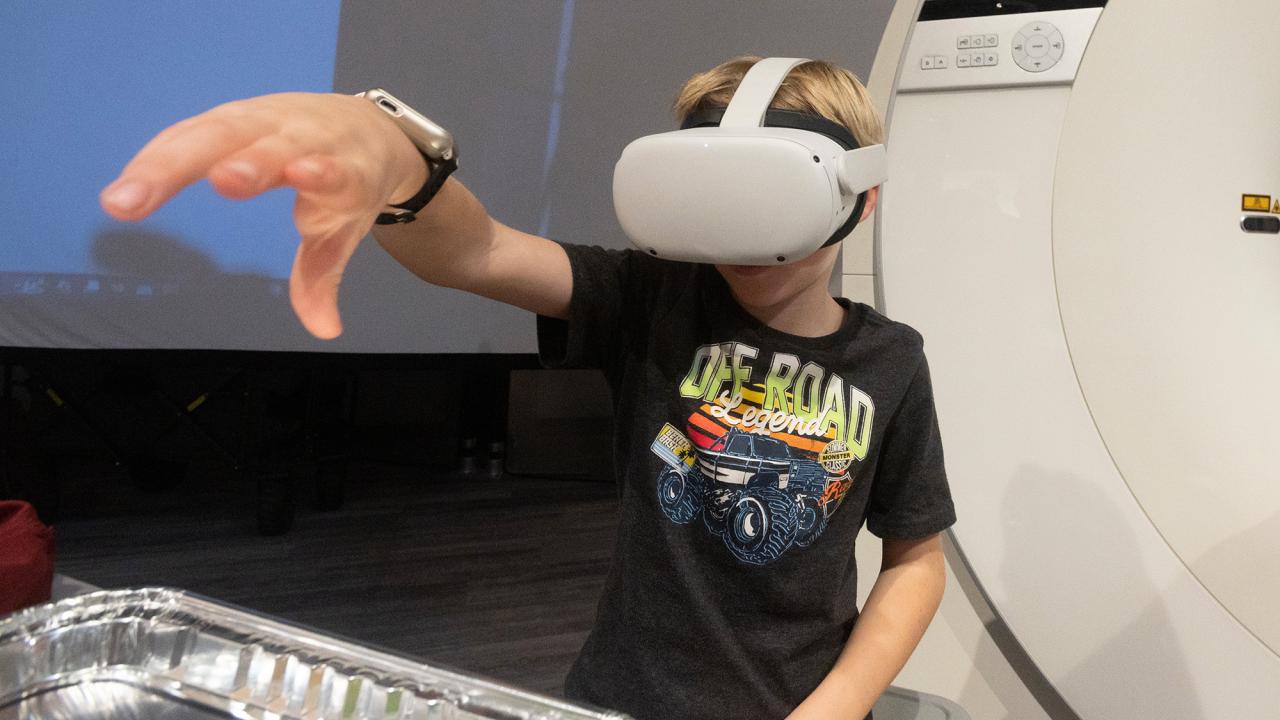
Unfold is a UC Davis podcast about science, innovation and discovery, unfolded through storytelling. We make complex topics relatable and reveal answers to questions you’ve always been curious about. We tackle big picture problems too, like how we’re going to feed a growing population, adapt to climate change and improve the health of people, animals and the planet.

Face to Face with Chancellor Gary May
Chancellor Gary S. May is used to being in the spotlight, and now he’s turning some attention on other Aggies making the world a better place. “I’m excited to be here talking with students, faculty and staff innovators about what they’re working on and how they’re making a difference in the real world.”

The Backdrop
The Backdrop podcast is a monthly interview program featuring conversations with UC Davis scholars and researchers working in the social sciences, humanities, arts and culture. Hosted by public radio veteran Soterios Johnson, the conversations feature new work and expertise on a trending topic in the news.